24/08/2020

Se desarrollan intensas pruebas contrarreloj para obtener la mejor vacuna anti-COVID-19. Dos organismos, uno del Reino Unido (Universidad de Oxford) y otro de China (CanSino Biologics) han logrado los resultados más prometedores hasta el momento. Un vector de adenovirus produce ambas vacunas (ChAdOx1 nCoV-19 y Adenovirus Type 5 Vector respectivamente) (de chimpancé y humano), un vector viral no replicante, que evita todo tipo de riesgos de infección en humanos. Sin embargo, es particularmente digno de mención que no solo estos dos estudios, sino la mayoría de los que están en desarrollo, utilizan la glicoproteína pico (S) del SARS-CoV-2 como protagonista principal de esta contrarreloj. Se han utilizado diferentes formatos para preparar/liberar esta proteína, como proteína recombinante purificada, como vector viral no replicante (como los dos intentos más avanzados descritos anteriormente) o como vacuna basada en ARN mensajero (ARNm).
Por otro lado, en una etapa temprana, esta proteína se destacó como el antígeno favorito para fines de diagnóstico, ya que esta proteína es esencial para la entrada del virus a las células huésped y probablemente la primera detectada por éstas como inmunógeno. Por este motivo, esta proteína forma parte de casi todos los test IVD para detección de COVID del mercado.
(...)
Ver más
08/04/2020
En este momento de pandemia, un punto muy crítico es saber si cada uno de nosotros estamos inmunizados y al mismo tiempo si contagiamos o no. Saber si las personas están inmunizadas contra el COVID-19 es una información imprescindible para poder deshacer este estado de cuarentena en el que estamos inmersos. Sólo las personas que hayan sido infectadas, pero ya no sean contagiosas, podrían volver a su vida normal, permitiendo un retorno progresivo a nuestra rutina anterior. Por ello, es fundamental analizar la carga de anticuerpos que presenta cada persona. Por tanto, la combinación de una RT-PCR + un test serológico rápido (para detección de anticuerpos) es lo que se debería imponer como requisito para volver a la normalidad. En primer lugar, una prueba serológica podría decirnos si tenemos anticuerpos contra el virus y sólo en el caso de que tengamos anticuerpos específicos en sangre será necesaria una confirmación por RT-PCR para asegurar que no somos infecciosos. Una IgG positiva con una IgM ya negativa, prácticamente no requeriría confirmación por RT-PCR, pero en ocasiones el virus o sus proteínas se mantienen en sangre por más tiempo. Así, existen diferentes posibilidades:
(...)
Ver más
24/02/2020
El 31 de diciembre de 2019, la OMS fue informada de un grupo de casos de neumonía de causa desconocida detectados en la ciudad de Wuhan, provincia china de Hubei. El 7 de enero las autoridades chinas identificaron la enfermedad por coronavirus (COVID-2019) como el virus causante. Como parte de la respuesta de la OMS al brote, se ha activado la I+D para acelerar los diagnósticos, las vacunas y las terapias para este nuevo coronavirus.
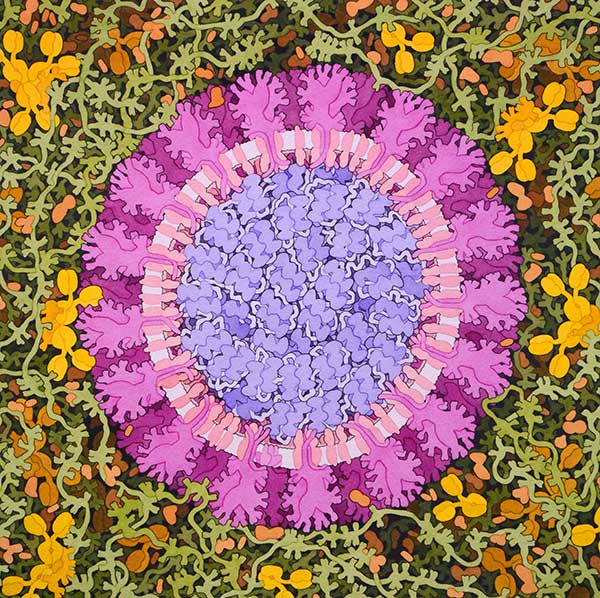

Figura 1. Coronavirus, 2020. Ilustración de David S. Goodsell, RCSB Protein Data Bank doi: 10.2210/rcsb_pdb/goodsell-gallery-019
(...)
Ver más
13/02/2020
Los antígenos nativos se extraen en su forma natural de la fuente correspondiente. En cambio, los antígenos recombinantes se fabrican artificialmente. Un método común para lograr esto es transformar el sistema de expresión heterólogo con un vector que exprese la proteína de interés, después de lo cual la proteína expresada se puede purificar del caldo de cultivo.
(...)
Ver más
09/05/2018
El citomegalovirus humano (HCMV) es endémico en poblaciones de todo el mundo, y entre el 50 y el 80 % de los adultos en los países desarrollados son seropositivos para este virus. Los ELISA comerciales para detectar IgG son muy sensibles y específicos. Sin embargo, existen varios problemas con respecto a la detección de IgM. Una de las dificultades a las que se ha enfrentado el laboratorio de diagnóstico durante los últimos 10 años es la falta de acuerdo entre las pruebas comerciales para la detección de IgM específica del CMV. Esta falta de acuerdo tiene su origen en las diferentes preparaciones virales utilizadas para detectar anticuerpos IgM contra el CMV. Rekom Biotech ha realizado el diseño de un nuevo CMV antígeno quimérico multi-epítopo recombinante (RAG0109), compuesto por diferentes determinantes antigénicos sensibles y específicos de algunas proteínas del CMV. Este antígeno quimérico multi-epítopo recombinante se probó en paralelo con otros biomarcadores de CMV, como las proteínas recombinantes de CMV de Rekom Biotech pp150 (RAG0091) y pp52 (RAG0090) y pp65 (RAG0016). El ensayo ELISA indirecto de IgM interno se llevó a cabo utilizando una batería de sueros prevalidados con la prueba ELISA comercial de captura de IgM de Vidas:
(...)
Ver más