8/24/2020

Intense time-trial is developed to obtain the best vaccine anti-COVID-19. Two organisms, one from the UK (Oxford University) and another from China (CanSino Biologics) have achieved the most promising results so far. An adenovirus vector produces both vaccines (ChAdOx1 nCoV-19 and Adenovirus Type 5 Vector respectively) (from chimp and human), a non-replicating viral vector, which avoids all type of infection risks in humans. However, it is particularly noteworthy, that not only these two studies but most of the developing ones are using the SARS-CoV-2 spike (S) glycoprotein as the leading player of this time trial. Different formats have been used to prepare/release this protein, as a purified recombinant protein, as a non-replicating viral vector (as the two more advanced attempts described above) or as a messenger RNA (mRNA)–based vaccine.
(...)
View more
4/8/2020
At this moment of the pandemic, a very critical point is to know if each of us is immunised and at the same time if is contagious or no. To know if people are immunised against COVID-19 is essential information to be able to undo this state of quarantine in which we are immersed. Only people who have been infected, but are already non-contagious, could return to their normal lives, allowing a progressive return to our previous routine. For this reason, it is essential to analyse the antibody load that each person presents. Therefore, the combined of an RT-PCR + a rapid serological test (for antibodies detection) is what should be imposed as a requirement to return to normality. First, a serological test could tell us if we have antibodies against the virus and only in the case we have specific antibodies in our blood, a confirmation by RT-PCR will be required to ensure that we are not infectious. A positive IgG with an already negative IgM, practically wouldn´t require confirmation by RT-PCR, but sometimes the virus or its proteins are maintained in blood for longer time. Thus, there exist different possibilities:
(...)
View more
2/24/2020
On 31 December 2019, WHO was informed of a cluster of cases of pneumonia of unknown cause detected in Wuhan City, Hubei Province of China. The coronavirus disease (COVID-2019) was identified as the causative virus by Chinese authorities on 7 January. As part of WHO’s response to the outbreak, the R&D has been activated to accelerate diagnostics, vaccines and therapeutics for this novel coronavirus.
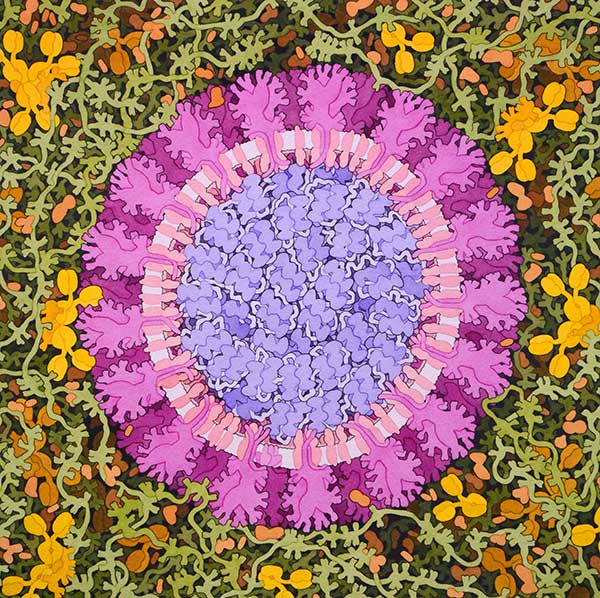

Figure 1. Coronavirus, 2020. Illustration by David S. Goodsell, RCSB Protein Data Bank doi: 10.2210/rcsb_pdb/goodsell-gallery-019
(...)
View more
2/13/2020
Native antigens are extracted in their natural form from the corresponding source. Recombinant antigens are instead manufactured artificially. A common method of achieving this is to transform the heterologous expression system with a vector expressing the protein of interest, following which the expressed protein can be purified from the culture broth.
(...)
View more
5/9/2018
Human cytomegalovirus (HCMV) is endemic to populations throughout the world, and 50-80% of the adults in developed nations are sero-positive for this virus. Commercial ELISAs for detecting IgG are very sensitive and specific. Nevertheless, there are several problems regarding IgM detection. One of the difficulties that the diagnostic laboratory has faced over the past 10 years is the lack of agreement between commercial tests for the detection of CMV-specific IgM. This lack of agreement has its roots in the different viral preparations used to detect IgM antibodies to CMV. Rekom Biotech has performed the design of a new CMV recombinant multi-epitope chimeric antigen (RAG0109), composed by different sensitive and specific antigenic determinants from some CMV proteins. This recombinant multi-epitope chimeric antigen was tested in parallel with other CMV bio-markers such as the Rekom Biotech CMV recombinant proteins pp150 (RAG0091) and pp52 (RAG0090) and pp65 (RAG0016). The in house indirect IgM ELISA assay was carried out by using a battery of pre-validated sera with the commercial ELISA capture IgM test from Vidas:
(...)
View more